Learning about antibodies
2026-03-25
Last updated: 2026-03-25
Checks: 7 0
Knit directory: muse/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20200712) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 1f3563d. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rproj.user/
Ignored: data/1M_neurons_filtered_gene_bc_matrices_h5.h5
Ignored: data/293t/
Ignored: data/293t_3t3_filtered_gene_bc_matrices.tar.gz
Ignored: data/293t_filtered_gene_bc_matrices.tar.gz
Ignored: data/5k_Human_Donor1_PBMC_3p_gem-x_5k_Human_Donor1_PBMC_3p_gem-x_count_sample_filtered_feature_bc_matrix.h5
Ignored: data/5k_Human_Donor2_PBMC_3p_gem-x_5k_Human_Donor2_PBMC_3p_gem-x_count_sample_filtered_feature_bc_matrix.h5
Ignored: data/5k_Human_Donor3_PBMC_3p_gem-x_5k_Human_Donor3_PBMC_3p_gem-x_count_sample_filtered_feature_bc_matrix.h5
Ignored: data/5k_Human_Donor4_PBMC_3p_gem-x_5k_Human_Donor4_PBMC_3p_gem-x_count_sample_filtered_feature_bc_matrix.h5
Ignored: data/97516b79-8d08-46a6-b329-5d0a25b0be98.h5ad
Ignored: data/Parent_SC3v3_Human_Glioblastoma_filtered_feature_bc_matrix.tar.gz
Ignored: data/brain_counts/
Ignored: data/cl.obo
Ignored: data/cl.owl
Ignored: data/jurkat/
Ignored: data/jurkat:293t_50:50_filtered_gene_bc_matrices.tar.gz
Ignored: data/jurkat_293t/
Ignored: data/jurkat_filtered_gene_bc_matrices.tar.gz
Ignored: data/pbmc20k/
Ignored: data/pbmc20k_seurat/
Ignored: data/pbmc3k.csv
Ignored: data/pbmc3k.csv.gz
Ignored: data/pbmc3k.h5ad
Ignored: data/pbmc3k/
Ignored: data/pbmc3k_bpcells_mat/
Ignored: data/pbmc3k_export.mtx
Ignored: data/pbmc3k_matrix.mtx
Ignored: data/pbmc3k_seurat.rds
Ignored: data/pbmc4k_filtered_gene_bc_matrices.tar.gz
Ignored: data/pbmc_1k_v3_filtered_feature_bc_matrix.h5
Ignored: data/pbmc_1k_v3_raw_feature_bc_matrix.h5
Ignored: data/refdata-gex-GRCh38-2020-A.tar.gz
Ignored: data/seurat_1m_neuron.rds
Ignored: data/t_3k_filtered_gene_bc_matrices.tar.gz
Ignored: r_packages_4.5.2/
Untracked files:
Untracked: .claude/
Untracked: CLAUDE.md
Untracked: analysis/.claude/
Untracked: analysis/aucc.Rmd
Untracked: analysis/bimodal.Rmd
Untracked: analysis/bioc.Rmd
Untracked: analysis/bioc_scrnaseq.Rmd
Untracked: analysis/chick_weight.Rmd
Untracked: analysis/likelihood.Rmd
Untracked: analysis/modelling.Rmd
Untracked: analysis/sampleqc.Rmd
Untracked: analysis/wordpress_readability.Rmd
Untracked: bpcells_matrix/
Untracked: data/Caenorhabditis_elegans.WBcel235.113.gtf.gz
Untracked: data/GCF_043380555.1-RS_2024_12_gene_ontology.gaf.gz
Untracked: data/SeuratObj.rds
Untracked: data/arab.rds
Untracked: data/astronomicalunit.csv
Untracked: data/davetang039sblog.WordPress.2026-02-12.xml
Untracked: data/femaleMiceWeights.csv
Untracked: data/lung_bcell.rds
Untracked: m3/
Untracked: women.json
Unstaged changes:
Modified: analysis/isoform_switch_analyzer.Rmd
Modified: analysis/linear_models.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/antibody.Rmd) and HTML
(docs/antibody.html) files. If you’ve configured a remote
Git repository (see ?wflow_git_remote), click on the
hyperlinks in the table below to view the files as they were in that
past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 1f3563d | Dave Tang | 2026-03-25 | Include additional background |
| html | f6051b2 | Dave Tang | 2023-06-27 | Build site. |
| Rmd | c90d5b7 | Dave Tang | 2023-06-27 | CDR sequences |
| html | b707032 | Dave Tang | 2023-06-26 | Build site. |
| Rmd | 9161c86 | Dave Tang | 2023-06-26 | Learning about antibodies |
Notes based on the study: Development of a humanized monoclonal antibody (MEDI-493) with potent in vitro and in vivo activity against respiratory syncytial virus.
This study describes the generation of a humanised monoclonal antibody, MEDI-493, that recognises a conserved neutralising epitope on the F glycoprotein of RSV. Broad neutralisation of a panel of 57 clinical isolates of the RSV A and B subtypes was demonstrated.
Antibody information
- UniProt Accession Number of Target Protein: P03420
- Alternative Name(s) of Target: F2; Fusion glycoprotein F0; Protein F; F gycoprotein; Human respiratory syncytial virus; humanized mAb 129
- Immunogen: The original antibody was generated by immunizing female BALB/c mice by primary intranasal infection with A2 strain of RSV.
- Specificity: This antibody recognizes the site A of the RSV F glycoprotein. This antibody binds the NSELLSLINDMPITNDQKKLMSNN epitope (PMID: 20098425).
Fusion glycoprotein sequence
Fusion glycoprotein F0 of Human respiratory syncytial virus A (strain A2).
>sp|P03420|FUS_HRSVA Fusion glycoprotein F0 OS=Human respiratory syncytial virus A (strain A2) OX=11259 GN=F PE=1 SV=1
MELLILKANAITTILTAVTFCFASGQNITEEFYQSTCSAVSKGYLSALRTGWYTSVITIE
LSNIKENKCNGTDAKVKLIKQELDKYKNAVTELQLLMQSTPPTNNRARRELPRFMNYTLN
NAKKTNVTLSKKRKRRFLGFLLGVGSAIASGVAVSKVLHLEGEVNKIKSALLSTNKAVVS
LSNGVSVLTSKVLDLKNYIDKQLLPIVNKQSCSISNIETVIEFQQKNNRLLEITREFSVN
AGVTTPVSTYMLT NSELLSLINDMPITNDQKKLMSNN VQIVRQQSYSIMSIIKEEVLAYV
VQLPLYGVIDTPCWKLHTSPLCTTNTKEGSNICLTRTDRGWYCDNAGSVSFFPQAETCKV
QSNRVFCDTMNSLTLPSEINLCNVDIFNPKYDCKIMTSKTDVSSSVITSLGAIVSCYGKT
KCTASNKNRGIIKTFSNGCDYVSNKGMDTVSVGNTLYYVNKQEGKSLYVKGEPIINFYDP
LVFPSDEFDASISQVNEKINQSLAFIRKSDELLHNVNAGKSTTNIMITTIIIVIIVILLS
LIAVGLLLYCKARSTPVTLSKDQLSGINNIAFSNAntibody sequence
Design of humanised VL and VH segments based on murine monoclonal antibody 1129. Human framework regions were derived from K102 for VL. VH framework (FR1) region was derived from Cor; remaining framework regions were derived from CE-1 sequence.
VL.
FR1 CDR1 FR2 CDR2
DIQMTQSPSTLSASVGDRVTITC KCQLSVGYMH WYQQKPGKAPKLLIY DTSKLAS
FR3 CDR3 FR4
GVPSRFSGSGSGTEFTLTISSLQPDDFATYYC FQGSGYPFT FGGGTKLEIKVH.
FR1 CDR1 FR2 CDR2
QVTLRESGPALVKPTQTLTLTCTFSGFSLS TSGMSVG WIRQPPGKALEWLA DIWWDDKKDYNPSLKS
FR3 CDR3 FR4
RLTISKDTSKNQVVLKVTNMDPADTATYYCAR SMITNWYFDV WGAGTTVTVSSBasics
Notes from Analysing antibody sequence for recombinant antibody expression.
- An antibody (a.k.a. immunoglobulin) is a large Y-shaped protein produced by B cells. They are used by the immune system to neutralise foreign objects.
- Fab region determines its antigen specificity (Fv region and CDR).
- Fc region determines the effects.
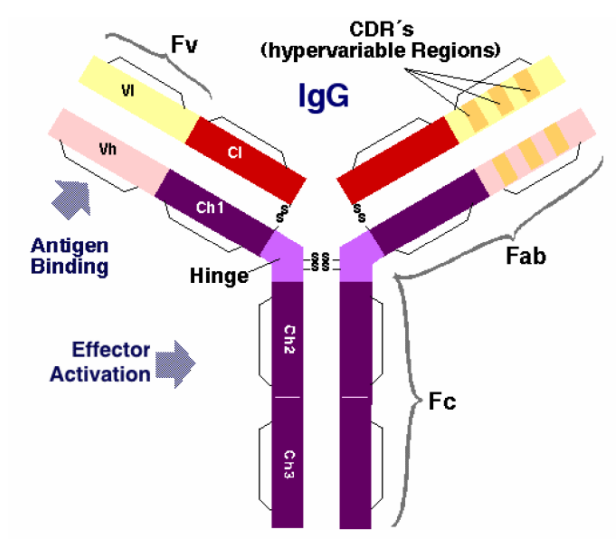
Antibody structure
Antibodies are composed of two identical heavy chains and two identical light chains, linked by disulfide bonds. Each chain consists of variable (V) and constant (C) domains:
- Heavy chain: One variable domain (VH) followed by three constant domains (CH1, CH2, CH3) for IgG/IgA/IgD, or four (CH1-CH4) for IgM/IgE.
- Light chain: One variable domain (VL) followed by one constant domain (CL).
Key structural regions:
- Fab fragment (Fragment antigen-binding): Contains VH-CH1 paired with VL-CL. Each antibody has two Fab arms.
- Fc fragment (Fragment crystallisable): Contains the CH2 and CH3 domains of both heavy chains. Mediates effector functions such as complement activation and binding to Fc receptors on immune cells.
- Hinge region: A flexible linker between CH1 and CH2 that allows the two Fab arms to move independently, facilitating bivalent antigen binding.
- Disulfide bonds: Inter-chain disulfide bonds in the hinge region link the two heavy chains; additional inter-chain bonds link each heavy chain to its paired light chain.
Each immunoglobulin domain adopts the immunoglobulin fold, a characteristic beta-sandwich structure of approximately 110 amino acids consisting of two beta-sheets stabilised by a conserved intra-domain disulfide bond.
Antigen-antibody interaction
- Each antibody (Ab) only binds to a specific antigen (Ag).
- Ab and Ag interact by spatial complementarity.
- Ab-Ag interaction are based on weak and non-covalent binding between
Ab and Ag:
- Electrostatic interactions
- Hydrogen bonds
- Van der Waals forces
- Hydrophobic interactions
- The hinge region allows better binding.
- Ab-Ag binding is reversible.

- Ab binding residues are mainly located in Complementarity Determining Regions (CDRs)
- 6 CDRs form the combining site: L1, L2, L3, H1, H2, and H3
- CDR-H3 and L3 have a distinctive role in antigen recognition.
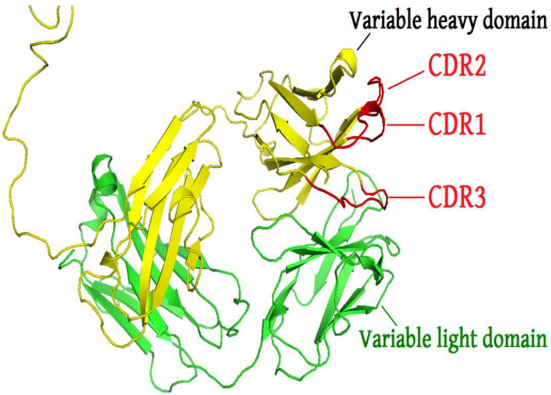
The framework region is a subdivision of the variable region (Fab) of the antibody. The variable region is composed of seven amino acid regions, four of which are framework regions (FR1-4) and three of which are hypervariable regions. The framework region makes up about 85% of the variable region. Located on the tips of the Y-shaped molecule, the framework regions are responsible for acting as a scaffold for the complementarity determining regions (CDR), also referred to as hypervariable regions, of the Fab. These CDRs are in direct contact with the antigen and are involved in binding antigen, while the framework regions support the binding of the CDR to the antigen and aid in maintaining the overall structure of the four variable domains on the antibody.
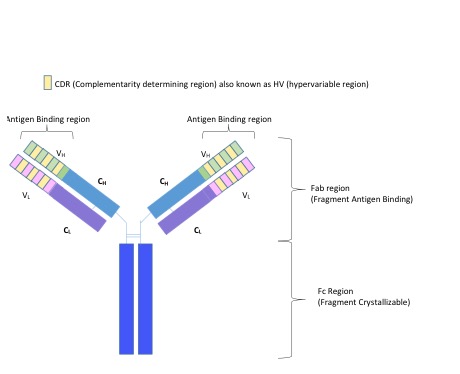
Antibody classes (isotypes)
There are five classes of antibodies in humans, distinguished by their heavy chain constant regions:
| Isotype | Heavy chain | Molecular form | Serum abundance | Key functions |
|---|---|---|---|---|
| IgG | \(\gamma\) (gamma) | Monomer | ~75% (most abundant) | Opsonisation, complement activation, ADCC, crosses placenta |
| IgA | \(\alpha\) (alpha) | Monomer or dimer | ~15% | Mucosal immunity, neutralisation at epithelial surfaces |
| IgM | \(\mu\) (mu) | Pentamer | ~10% | First antibody in primary immune response, strong complement activation |
| IgD | \(\delta\) (delta) | Monomer | <1% | B cell receptor signalling |
| IgE | \(\epsilon\) (epsilon) | Monomer | <0.01% | Allergic responses, anti-parasitic immunity |
IgG subclasses
IgG is further divided into four subclasses in humans with different effector properties:
| Subclass | Abundance | Complement | Fc\(\gamma\)R binding | Half-life | Typical targets |
|---|---|---|---|---|---|
| IgG1 | ~60% | +++ | Strong | ~21 days | Protein antigens |
| IgG2 | ~25% | + | Weak | ~21 days | Polysaccharide antigens |
| IgG3 | ~5% | +++ | Strong | ~7 days | Protein antigens |
| IgG4 | ~5% | - | Intermediate | ~21 days | Repeated/chronic antigen exposure |
Most therapeutic monoclonal antibodies, including palivizumab (MEDI-493), use an IgG1 framework due to its strong effector functions and long serum half-life.
Light chain types
There are two types of light chains:
- Kappa (\(\kappa\)): Encoded on chromosome 2 in humans. ~60% of human antibodies use kappa light chains.
- Lambda (\(\lambda\)): Encoded on chromosome 22 in humans. ~40% of human antibodies use lambda light chains.
A given antibody molecule always contains two identical light chains (either both kappa or both lambda, never mixed). The choice of light chain type does not affect antigen specificity or effector function.
Identifying CDRs by sequence
CDR-L1
- Start - approximately residue 24
- Residue before - always a Cys
- Residue after - always a Trp. Typically Trp-Tyr-Gln, but also Trp-Leu-Gln, Trp-Phe-Gln, Trp-Tyr-Leu
- Length - 10 to 17 residues
CDR-L2
- Start - always 16 residues after the end of L1
- Residue before - generally Ile-Tyr, but also, Val-Tyr, Ile-Lys, Ile-Phe
- Length - always 7 residues (except NEW [7FAB], which has a deletion in this region.)
CDR-L3
- Start - always 33 residues after end of L2 (except NEW [7FAB], which has the deletion at the end of CDR-L2)
- Residue before - always Cys
- Residue after - always Phe-Gly-XXX-Gly
- Length - 7 to 11 residues
CDR-H1
- Start - approximately residue 26 (always 4 after a Cys)
- Residue before - always Cys-XXX-XXX-XXX
- Residue after - always a Trp. Typically Trp-Val, but also, Trp-Ile, Trp-Ala
- Length - 10 to 12 residues
CDR-H2
- Start - always 15 residues after the end of Kabat / AbM definition of CDR-H1
- Residue before - typically Leu-Glu-Trp-Ile-Gly, but a number of variations
- Residue after - Lys/Arg-Leu/Ile/Val/Phe/Thr/Ala-Thr/Ser/Ile/Ala
- Length - Kabat definition 16 to 19 residues
CDR-H3
- Start - always 33 residues after end of CHR-H2 (always 2 after a Cys)
- Residue before - always Cys-XXX-XXX (typically Cys-Ala-Arg)
- Residue after - always Trp-Gly-XXX-Gly
- Length - 3 to 25 residues
Leader sequence-FR1-CDR1-FR2-CDR2-FR3-CDR3-FR4-Constant region-Stop codon
Generating antibody diversity
V(D)J recombination
Antibody diversity is generated primarily through V(D)J recombination, a somatic recombination process that occurs during B cell development in the bone marrow.
The immunoglobulin heavy chain locus contains multiple gene segments:
- V (Variable): ~40 functional segments
- D (Diversity): ~25 functional segments
- J (Joining): 6 functional segments
The light chain loci (kappa and lambda) contain V and J segments but lack D segments.
During B cell development, the RAG1/RAG2 recombinase enzymes randomly select and join:
- One D to one J segment
- One V to the DJ segment (for heavy chains)
- One V to one J segment (for light chains)
Additional diversity is introduced by:
- Junctional diversity: Imprecise joining at V-D and D-J junctions, including addition of P-nucleotides and N-nucleotides by terminal deoxynucleotidyl transferase (TdT).
- Combinatorial diversity: Random pairing of heavy and light chains.
Together, these mechanisms generate an estimated theoretical diversity of >10^11 unique antibodies.
Somatic hypermutation and affinity maturation
After initial antigen encounter, B cells in germinal centres undergo somatic hypermutation (SHM), introducing point mutations at a high rate (~10^-3 per base pair per cell division) in the variable regions of immunoglobulin genes. This process is mediated by activation-induced cytidine deaminase (AID).
B cells with mutations that improve antigen binding are positively selected through interaction with follicular dendritic cells and T follicular helper cells, while those with reduced affinity undergo apoptosis. This iterative process of mutation and selection is called affinity maturation and results in antibodies with progressively higher affinity for their target antigen.
Class switch recombination (CSR) can also occur in germinal centres, allowing B cells to switch from producing IgM to other isotypes (IgG, IgA, IgE) while retaining the same antigen specificity.
Monoclonal vs polyclonal antibodies
Polyclonal antibodies: A mixture of antibodies produced by different B cell clones, recognising multiple epitopes on the same antigen. Generated by immunising an animal and collecting serum. Advantages: high overall avidity, tolerance to minor antigen changes. Disadvantages: batch-to-batch variability, limited supply.
Monoclonal antibodies (mAbs): Antibodies produced by a single B cell clone, recognising a single epitope. First generated using hybridoma technology (Köhler and Milstein, 1975): B cells from an immunised mouse are fused with myeloma cells to create immortal antibody-producing cell lines. Advantages: high specificity, unlimited reproducible supply, consistent properties. Disadvantages: more complex to generate, may have limited recognition of conformational variants.
Antibody engineering
Therapeutic antibodies have evolved through several generations of engineering to reduce immunogenicity in humans:
- Murine antibodies (-omab): Entirely derived from mouse. Highly immunogenic in humans, eliciting human anti-mouse antibody (HAMA) responses.
- Chimeric antibodies (-ximab): Mouse variable regions joined to human constant regions. ~33% murine sequence. Example: rituximab.
- Humanised antibodies (-zumab): Only the CDRs from the mouse antibody are grafted onto a human framework. ~5-10% murine sequence. Key framework residues from the original mouse antibody may be retained (back-mutations) to preserve CDR conformation and binding affinity. Example: palivizumab, trastuzumab.
- Fully human antibodies (-umab): Entirely human sequence, generated using transgenic mice with human immunoglobulin loci or phage display libraries. Example: adalimumab.
Palivizumab (MEDI-493) described in this study is a humanised antibody, with CDRs from murine mAb 1129 grafted onto human framework regions (K102 for VL; Cor and CE-1 for VH).
Note: The -mab suffix naming convention was updated by the WHO/INN in 2021. Newer antibodies may not follow these suffixes.
Antibody numbering schemes
Several numbering schemes exist for consistently numbering antibody residues across different sequences, which is critical for comparing CDR boundaries and framework regions:
- Kabat numbering: Based on sequence variability analysis of known antibody sequences. The most widely used scheme historically. CDR definitions are based on sequence variability. Used in the CDR identification rules described above.
- Chothia numbering: Based on structural analysis of antibody crystal structures. CDR definitions reflect the actual loop structures. The CDR-H1 definition differs significantly from Kabat (Chothia starts 5 residues earlier and ends 1 residue earlier).
- IMGT numbering: Developed by the International ImMunoGeneTics information system. Uses a standardised framework with a fixed number of positions per region, inserting gaps to maintain alignment. Provides the most consistent cross-species comparison.
- AHo numbering: Based on structural alignment of all immunoglobulin and T-cell receptor domains.
| Scheme | CDR-L1 | CDR-L2 | CDR-L3 | CDR-H1 | CDR-H2 | CDR-H3 |
|---|---|---|---|---|---|---|
| Kabat | 24-34 | 50-56 | 89-97 | 31-35B | 50-65 | 95-102 |
| Chothia | 24-34 | 50-56 | 89-97 | 26-32 | 52-56 | 95-102 |
| IMGT | 27-38 | 56-65 | 105-117 | 27-38 | 56-65 | 105-117 |
Online tools such as ANARCI and AbNum can automatically number antibody sequences using these schemes.
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: Etc/UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.0
[5] purrr_1.2.1 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[9] ggplot2_4.0.2 tidyverse_2.0.0 workflowr_1.7.2
loaded via a namespace (and not attached):
[1] sass_0.4.10 generics_0.1.4 stringi_1.8.7 hms_1.1.4
[5] digest_0.6.39 magrittr_2.0.4 timechange_0.4.0 evaluate_1.0.5
[9] grid_4.5.2 RColorBrewer_1.1-3 fastmap_1.2.0 rprojroot_2.1.1
[13] jsonlite_2.0.0 processx_3.8.6 whisker_0.4.1 ps_1.9.1
[17] promises_1.5.0 httr_1.4.8 scales_1.4.0 jquerylib_0.1.4
[21] cli_3.6.5 rlang_1.1.7 withr_3.0.2 cachem_1.1.0
[25] yaml_2.3.12 otel_0.2.0 tools_4.5.2 tzdb_0.5.0
[29] httpuv_1.6.17 vctrs_0.7.2 R6_2.6.1 lifecycle_1.0.5
[33] git2r_0.36.2 fs_2.0.0 pkgconfig_2.0.3 callr_3.7.6
[37] pillar_1.11.1 bslib_0.10.0 later_1.4.8 gtable_0.3.6
[41] glue_1.8.0 Rcpp_1.1.1 xfun_0.57 tidyselect_1.2.1
[45] rstudioapi_0.18.0 knitr_1.51 farver_2.1.2 htmltools_0.5.9
[49] rmarkdown_2.30 compiler_4.5.2 getPass_0.2-4 S7_0.2.1